3D Interpolation¶
Once we register consecutive 2D slices the next step is to think of creating 3D volumes. This is especially relevant for data from the Allen Brain Institute that come from experiments with a very specific design.
There are multiple ways how to perform this interpolation and some of these are contained in the atlalign.volume
module:
CoronalInterpolator
CoronalInterpolator¶
The CoronalInterpolator turns the entire problem into a 1D function interpolation. See below the sketch:
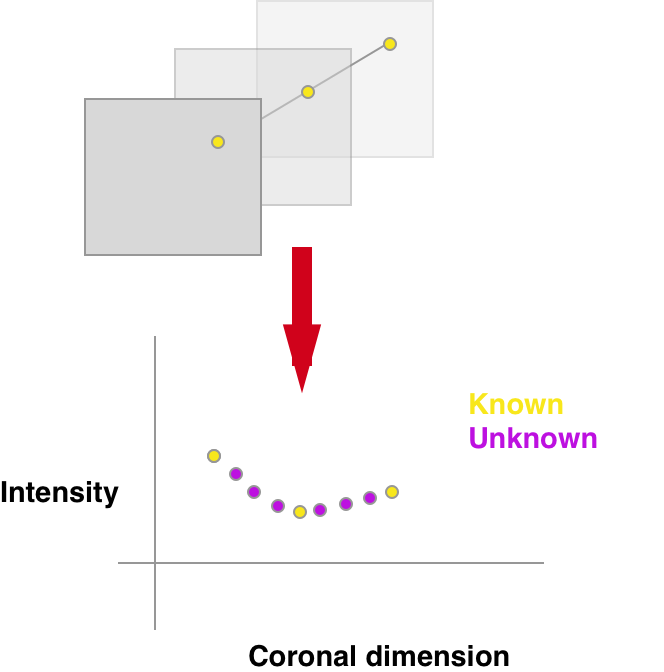
For each pixel separately one interpolates over missing sections based purely on the corresponding pixels in all of the existing sections.
import numpy as np
from atlalign.volume import CoronalInterpolator, GappedVolume
n_sections = 55
shape = (30, 40)
sn = np.random.choice(np.arange(527), size=n_sections, replace=False)
imgs = np.random.random((n_sections, *shape))
gv = GappedVolume(sn, imgs)
ci = CoronalInterpolator()
final_volume = ci.interpolate(gv)
print(final_volume.shape) # (528, shape[0], shape[1])